Literature
Question:
How do I identify HCPs with low molecular weight?
I work on a drug substance protein with a mass of 15 kDa. I would expect small host cell protein impurities to co-purify with the drug substance, but HCP analysis by 2D gels, Western blots, and HCP ELISA kits shows only a few low Mw HCPs. Is there an LC-MS/MS method for identifying host cell proteins with low molecular weight (small HCPs below 20 kDa)?
Answer:
Yes, there are some excellent mass spectrometry methods for identifying HCPs with small MW. The methods are based on trypsin digestion, HPLC separation of the peptides, and MS/MS peptide analysis to identify the proteins [1, 2].
I find that the two best LC-MS/MS methods we have at Alphalyse for small MW HCPs are:
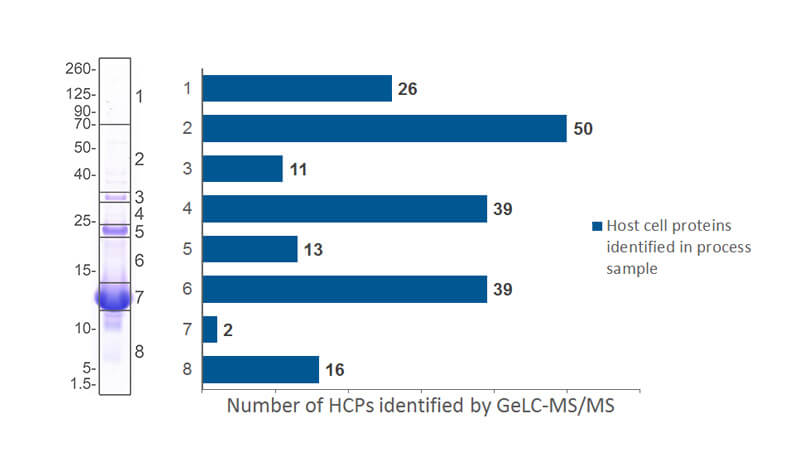
- GeLC-MS/MS, where we fractionate your drug substance sample by Mw in 1D SDS PAGE and hereafter identify HCPs by nanoLC-MS/MS.
- LC-MS/MS with SWATH data acquisition of your drug sample.
Both methods are optimized to detect the very low concentrations of HCPs in a high content of drug substance. Typically from 1-1000 parts per million (ppm).
Moreover, we use a database search algorithm that allows the identification of proteins with a good matching of only 1-2 peptides. We do this not to exclude small HCPs with only very few tryptic peptides.
Common reasons why small HCPs are challenging to detect
Small HCPs are often difficult to detect by other methods for several reasons:
- Small Mw proteins don’t stain well by total protein stains in 1D and 2D gels.
The stain binds to specific amino acids, e.g., Coomassie blue binds to lysine and arginine residues. Since small proteins have a lower total number of these residues, they don’t stain as well as larger ones [3-5]. - Small Mw proteins are poor immunogens, and raising suitable antibodies against small proteins is challenging.
Peptides and small proteins are often less immunogenic than larger proteins. The immunized animals, therefore, fail to produce antibodies against low Mw HCPs. Hence, the generated HCP antibodies fail to detect any small HCPs in the ELISA and the corresponding Western blot [3-5].
Increase the likelihood of detecting small MW HCPs
So, how does one achieve the best identification results? In my experience, it is a good idea to:
- Prepare the HCP antibodies with specific attention to the low Mw protein fraction.
Immunization with pools of host cell proteins only containing low Mw HCPs will increase the likelihood of an immunogenic response against these. It can be achieved by size-fractionating the null cell sample proteins before immunization. With this, you avoid competition with the more immunogenic high Mw HCPs [5]. - Finally, ensure that you analyze the samples using an LC-MS/MS method specifically developed to detect HCP levels of low Mw proteins.
Small HCPs only contain very few tryptic peptides, and the mass spectrometer will not detect all of them. Hence, small proteins are excluded if you use a Hi3 algorithm requiring at least three peptides to identify the proteins. Therefore, you must use a protein identification algorithm to identify proteins only based on one or two peptides [5].
Sources for more information on the analysis of small MW HCPs
Rikke Lund has written a very comprehensive application note on the subject. You will find the application note along with poster by clicking the links or on the Alphalyse website.
References
[1] Guiochon et al.: “Separation science is the key to successful biopharmaceuticals,” Journal of Chromatography A, 2011
[2] Tscheliessnig et al.: “Host cell protein analysis in therapeutic protein bioprocessing – methods and applications,“ Biotechnology Journal, 2013
[3] Dimitrov, Dimiter: “Therapeutic proteins,” Methods in Molecular Biology, 2012
[4] Wang et al.: “Host Cell Proteins in Biologics Development: Identification, Quantitation and Risk Assessment,” Biotechnology and Bioengineering, 2009
[5] Bracewell et al.: “The Future of Host Cell Protein (HCP) Identification During Process Development and Manufacturing Linked to a Risk-Based Management for Their Control,” Biotechnology and Bioengineering, 2015
Talk to us
Whatever protein-related challenge or question you may have, we would love to help. Our experts can help you decide on the best analytical approach for your project by email or online meeting - providing advice without obligation.
