Literature
Question:
Does our generic ELISA kit have sufficient coverage?
My company uses a commercial HCP ELISA kit to analyze clinical phase 1 material host cell protein. We hear that we must develop a process-specific ELISA for clinical phases 2 and 3. Developing a process-specific ELISA could take two years, cost a fortune, and might not even have good coverage.
Is there a way to ensure that we pick a HEK293 HCP ELISA kit with high coverage? And thus increase the chances of using it for approval of clinical-stage trials.
Answer:
The short answer is – yes.
Many people still believe clinical trials require the tiresome development of process-specific ELISA antibodies – and is unavoidable. It is probably because commercial kits sometimes have a low HCP coverage*. (*the percentage of host cell proteins (HCPs) recognized by the antibodies [1]).
A suitable assay could be any ELISA documented to cover enough HCPs
However, recent recommendations from government officials at FDA and EMA suggest otherwise. They only say that the chosen HCP assay should have sufficient coverage and demonstrated ability to detect most HCPs in the product [2].
Based on this, a suitable HCP coverage assay could be any ELISA documented to cover enough HCPs. And thus, even a commercial HCP ELISA with proper validation of its coverage [3], like CHO, E. coli, and HEK 293.
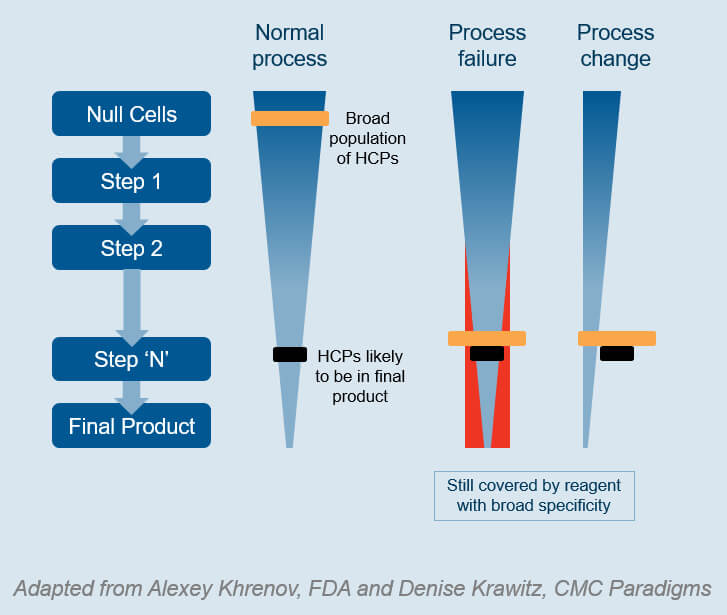
Now, let’s compare methods for evaluation of HCP ELISA kits:
Typically, you evaluate the coverage by protein spot counting, using 2D SDS-PAGE and Western Blotting, or Immunoaffinity binding and 1D/2D SDS-PAGE [3].
This method’s disadvantages are long: It includes high variability, low reproducibility, and a high risk of not detecting abundant HCPs of low MW. Furthermore, the HCP antigens denature during 2D PAGE analysis. In contrast, the method detects antigens in their native form in ELISA [4].
So what if a biopharmaceutical company wishes to skip process-specific antibody development? Then it needs a better coverage analysis to determine the precise coverage percent of the ELISA for specific HCPs. It enables comparing and selecting the ELISA with the best coverage of all HCPs, whether commercial or process-specific [3, 4, 5].
The ideal coverage method includes a complete list of HCPs covered by the assay to select the best HCP ELISA [3, 4].
Fortunately, a method that can achieve this is now available and described below:
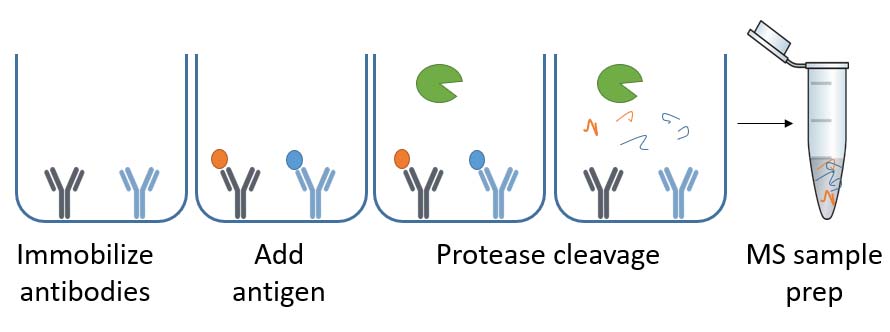
What is ELISA-based immunocapture?
This method first immobilizes the antibodies to the 96-well plate, like the sandwich ELISA. Subsequently, process samples are added, and antigens bind to their respective antibodies – if the HCP ELISA antibody kit contains them. Finally, the process digests the bound antigens with proteolytic enzymes before mass spectrometry (MS) sample prep and LC-MS/MS analysis.
Information-dependent acquisition (IDA) liquid chromatography (LC) tandem mass spectrometry (MS/MS) can find the proteins covered by the HCP ELISA. This method is denoted IDA LC-MS/MS or SWATH LC-MS/MS and uses the Sciex TripleTOF 6600 system. It also includes searches of relevant protein databases.
The analysis provides you with:
- A list of proteins recognized by the ELISA antibodies
- A list of all proteins in mock or process sample
- The HCP coverage percentage
- The specific coverage of HCPs in the drug product
What else should you know about evaluating the coverage of HCP ELISA kits?
The new analysis was first introduced at BEBPA 2019 conference, the largest HCP conference in the world. This poster describes the method in more detail:
More information about the coverage analysis is available on the Alphalyse website.
References
[1] Bracewell et al.: “The Future of Host Cell Protein (HCP) Identification During Process Development and Manufacturing Linked to a Risk-Based Management for Their Control,” Biotechnology and Bioengineering, 2015
[2] The United States Pharmacopeia: “Residual Host Cell Protein Measurement in Biopharmaceuticals,” USP 39 – NF 34, 2016
[3] Zhu-Shimoni et al.: “Host Cell Protein Testing by ELISAs and the Use of Orthogonal Methods,” Biotechnology and Bioengineering, 2014
[4] Wang et al.: “Host Cell Proteins in Biologics Development: Identification, Quantitation and Risk Assessment,” Biotechnology and Bioengineering, 2009
Talk to us
Whatever protein-related challenge or question you may have, we would love to help. Our experts can help you decide on the best analytical approach for your project by email or online meeting - providing advice without obligation.
